Docking in Flare™ is one of the platform’s most popular functions, providing its users with detailed feedback on new molecule designs, high enrichment in virtual screening, and excellent pose prediction. Flare V7 now includes an expanded choice and more flexibility within its available docking workflows, in support of more accurate protein-ligand binding predictions. In this article, we will be highlighting the new covalent warheads which are now available in Flare V7, as well as showing you what can be done with its covalent docking feature.
What is a covalent warhead?
Although ‘warhead’ sounds aggressive, a covalent warhead is simply a functional group or substructure within a small molecule that reacts with a reactive residue in a protein. Sometimes, it’s present in a natural product, such as the acetyl group in aspirin, and other times in a synthetic drug, such as the nitrile in Nirmatrelvir.1
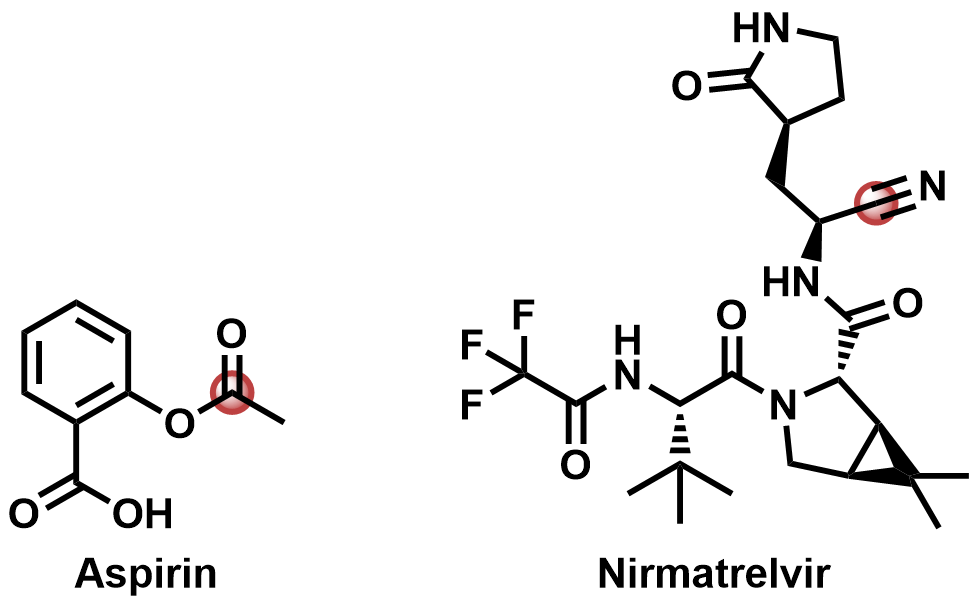 Figure 1. Aspirin (left) with its reactive acetyl group highlighted; Nirmatrelvir (right) with its reactive nitrile highlighted.
Figure 1. Aspirin (left) with its reactive acetyl group highlighted; Nirmatrelvir (right) with its reactive nitrile highlighted.
A covalent warhead forms a covalent bond, either reversibly or irreversibly, with a nucleophilic residue found in the binding site of the target. Irreversibly reactive covalent warheads, such as Michael acceptors or epoxides, permanently modify the protein in question. Conversely, reversibly reactive covalent warheads, such as nitriles or boronic acids, can eventually leave the active site.2
When should covalent docking be used, and what is available in Flare?
Simply put, you should use covalent docking when you have a ligand containing a reactive functional group that could form a covalent bond with a nucleophilic residue in your protein.
Currently, we have 20 available covalent warheads recognized by Flare. Newly added are the aldehyde, aryl-activated ketone, and pentafluorobenzenesulfonamide (PFBS).
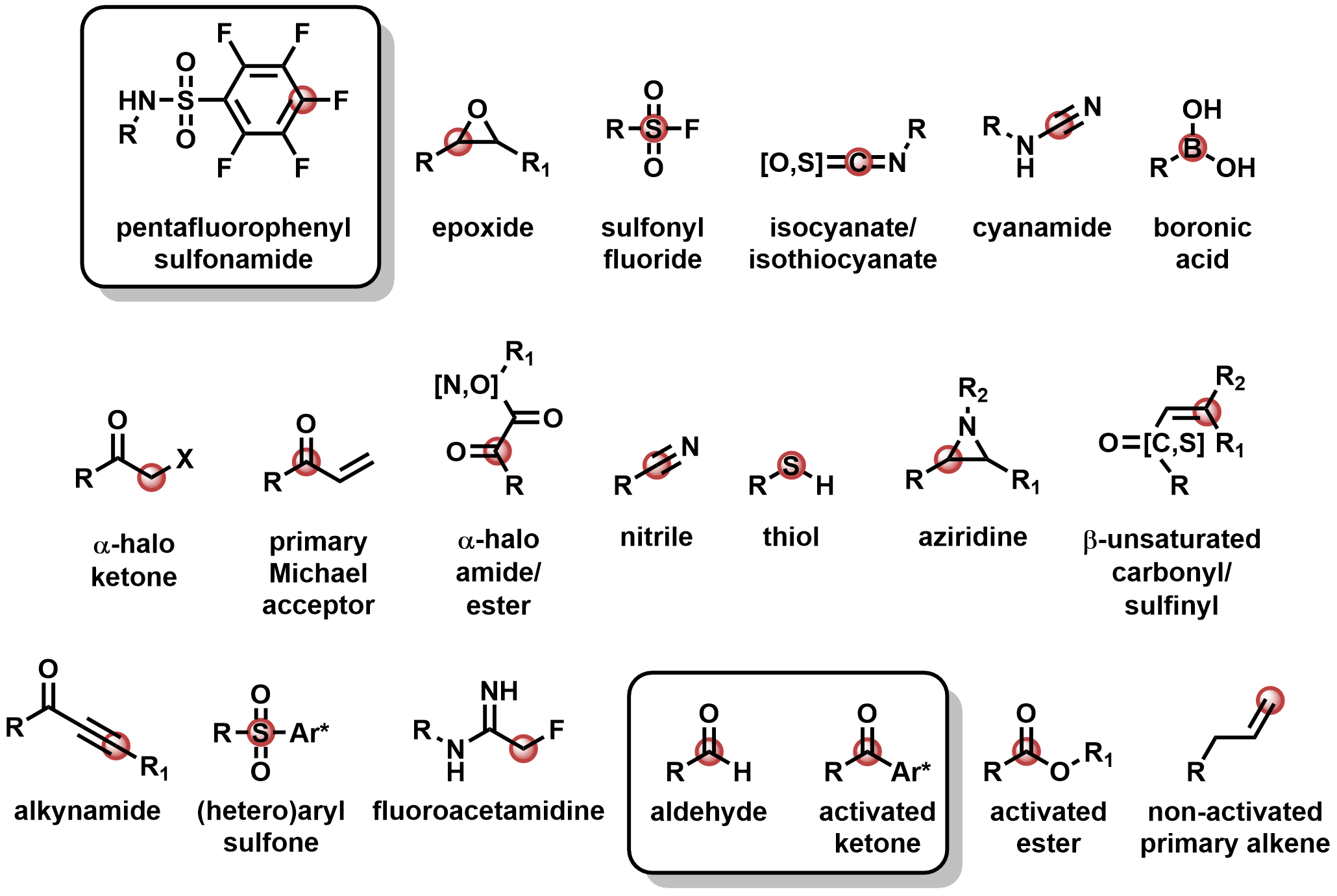 Figure 2. Covalent warheads currently implemented within Flare, organized by order of priority as recognized by the program. Newly added warheads are highlighted in the box: the aldehyde, activated ketone, and pentafluorophenyl sulfonamide.
Figure 2. Covalent warheads currently implemented within Flare, organized by order of priority as recognized by the program. Newly added warheads are highlighted in the box: the aldehyde, activated ketone, and pentafluorophenyl sulfonamide.
How to begin a covalent docking experiment in Flare
Running a covalent docking experiment in Flare is very similar to a ‘Normal’ docking workflow. Once you have prepared your protein and extracted your reference ligand, select the ligands you wish to dock covalently. Under the ‘3D Pose’ tab, go to ‘Dock > Covalent’. Define your grid as you normally would, and then identify the covalent residue with the dropdown options next to ‘Residue’.
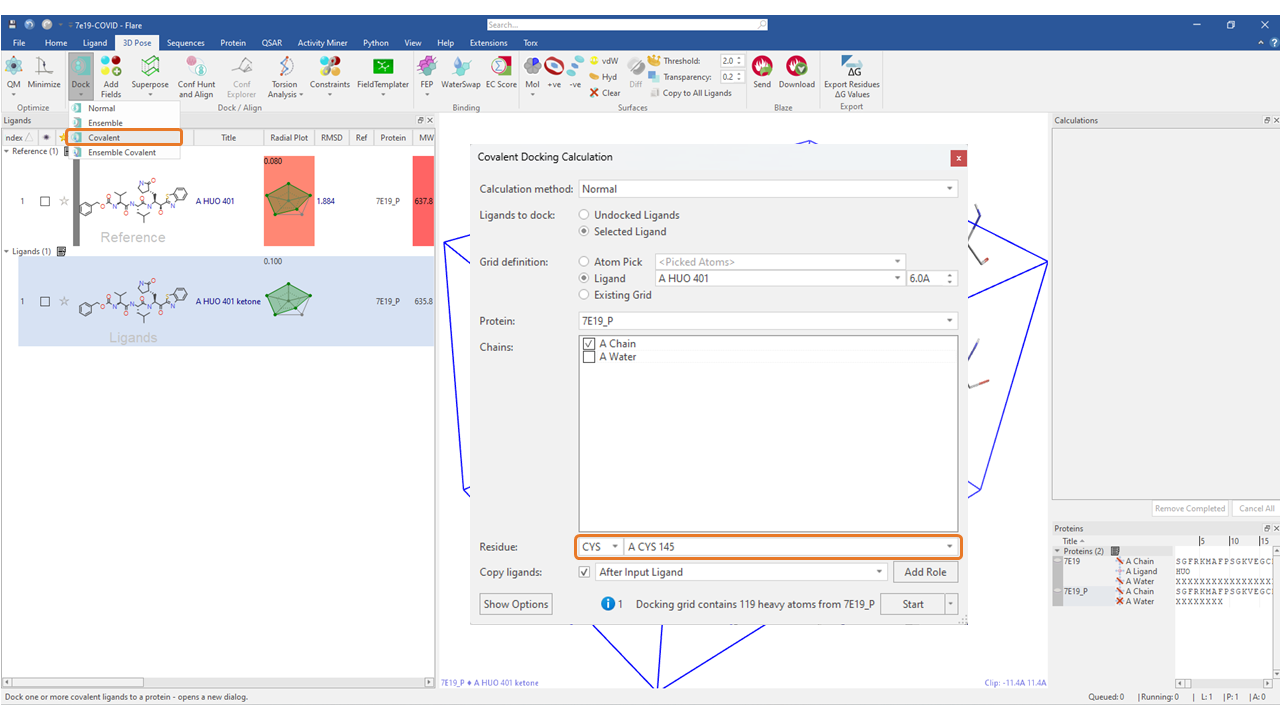 Figure 3. The Covalent docking workflow in Flare.
Figure 3. The Covalent docking workflow in Flare.
An example of the aldehyde warhead
The aldehyde is a very common warhead for studying the kinetics of protease enzymes, as the resultant hemiacetal forms a tetrahedral complex with the catalytic residue and surrounding stabilizing residues, mimicking the transition state of the native peptide substrates. For example, prolyl oligopeptidase (POP) is a protease enzyme implicated in Alzheimer’s disease and other neurodegenerative diseases. The enzymatic mechanism associated with these diseases involves the cleavage of native peptides by a catalytic serine residue in its active site. In their work, the authors used Z-Pro-Prolinal (ZPP) inhibitor, a dipeptide derivative containing an aldehyde covalent warhead.3 Here, ZPP has been docked into POP using Flare’s covalent docking workflow (Figure 4).
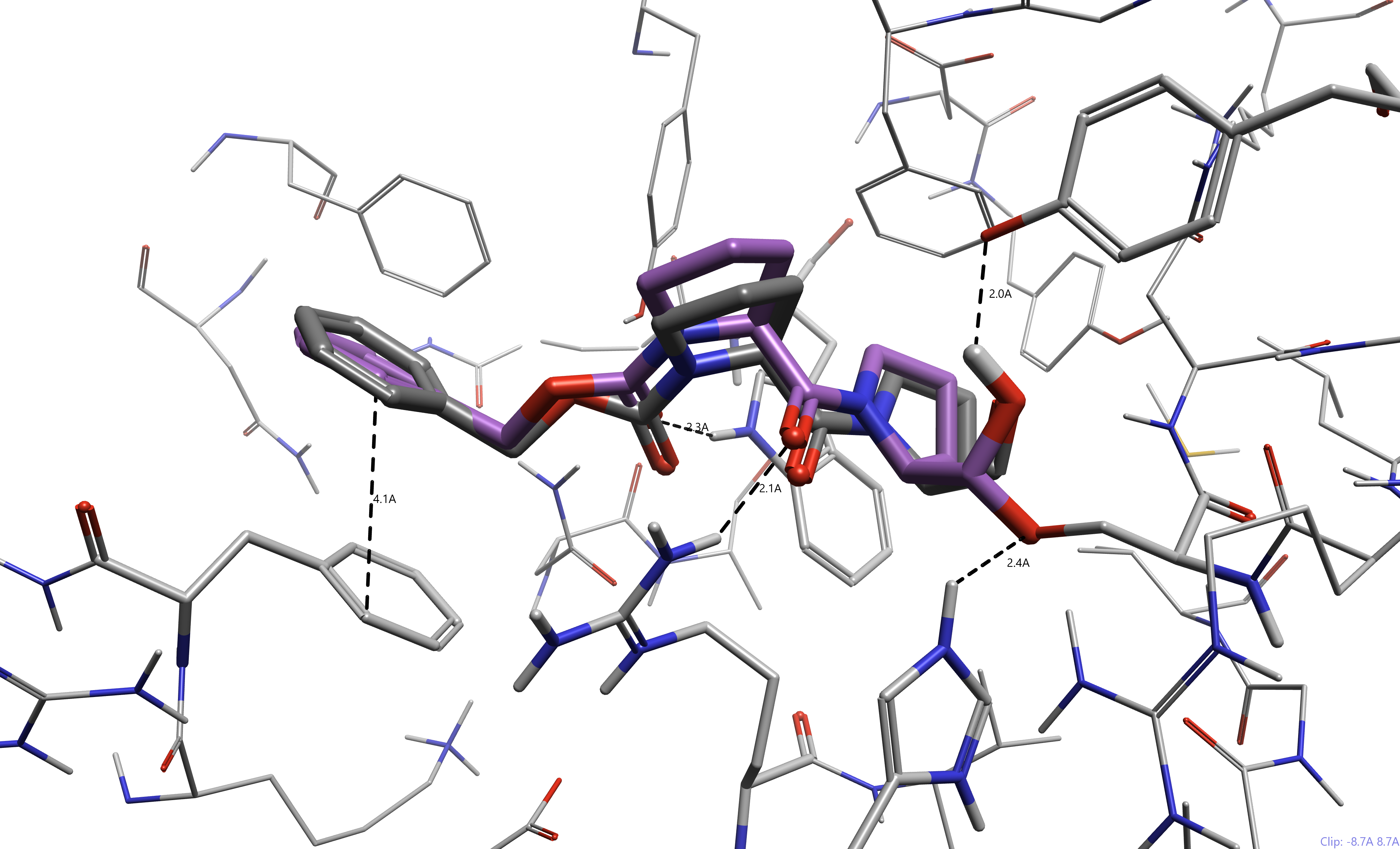 Figure 4. Z-Pro-Prolinal docked to prolyl oligopeptidase (pdb 7VGC). The docked ligand is represented in violet, while the co-crystallized ligand is represented in dark gray. Stabilizing interactions shown in black, catalytic serine highlighted.
Figure 4. Z-Pro-Prolinal docked to prolyl oligopeptidase (pdb 7VGC). The docked ligand is represented in violet, while the co-crystallized ligand is represented in dark gray. Stabilizing interactions shown in black, catalytic serine highlighted.
You can see the docking pose is very similar to the binding mode of the ligand in the crystal structure (RMSD = 0.8 Å). Both bound structures are participating in the same key non-covalent stabilization interactions, as well as forming the resultant hemiacetal with the catalytic serine. Additionally, the highly flexible nature of this peptidomimetic drug is docked very close to the original crystal structure, further highlighting the accuracy of the Flare docking feature.
An example of the activated ketone warhead
The activated ketone warhead is another common moiety found in protease inhibitors.4-5 In recent literature, it has been utilized to target SARS-CoV2 3CL Pro to block viral replication.6 The activated ketone reacts similarly to the aldehyde, forming a hemiacetal complex with the nucleophilic protein residue. In Figure 5, a recently published inhibitor YH-53 by Konno et al. (pdb 7E19)6 has been docked.
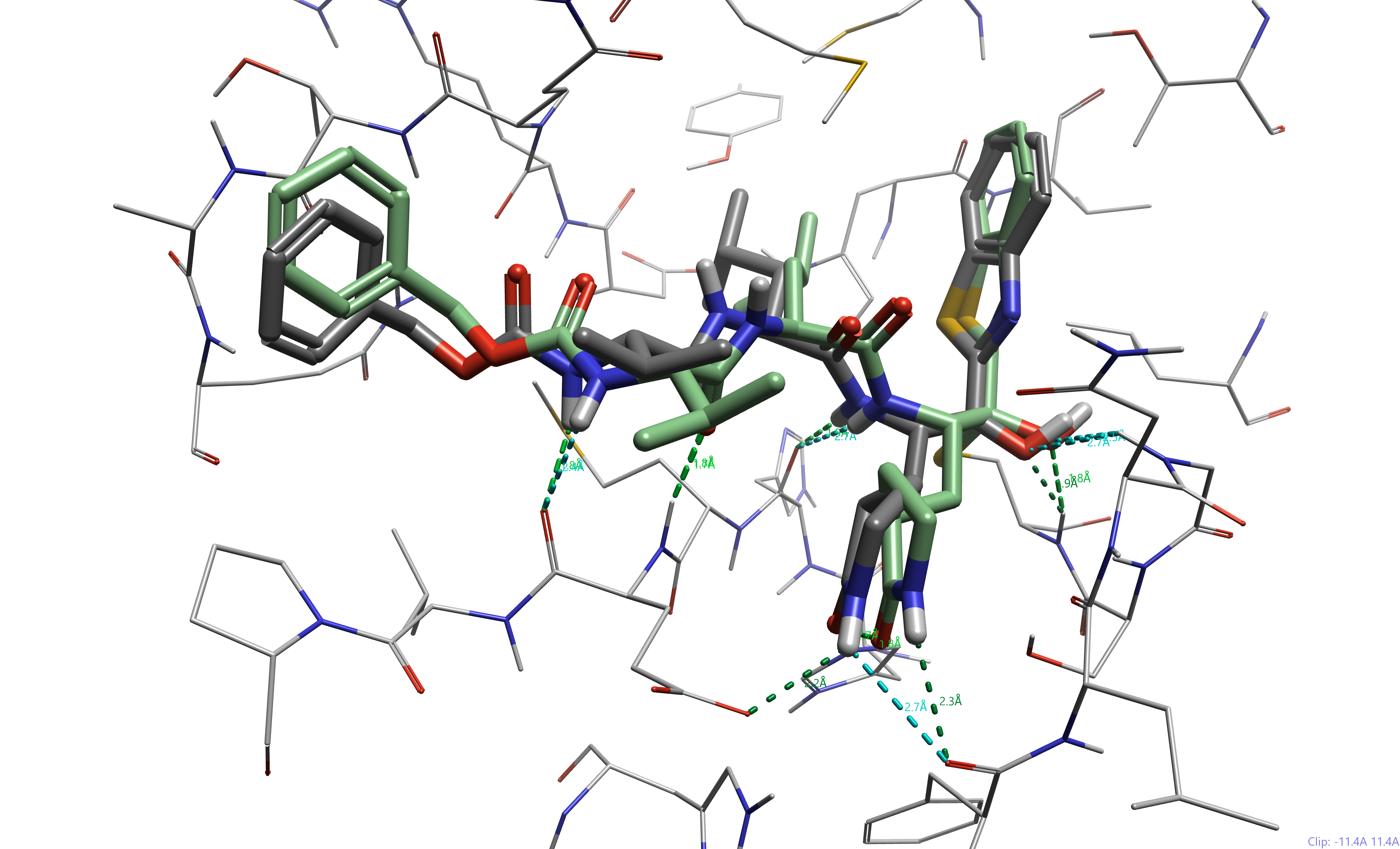 Figure 5. Compound YH-53 docked to SARS-CoV2 3CL Pro (pdb 7E19). The docked ligand is represented in green, while the co-crystallized ligand is represented in dark gray. Hydrogen bond contacts are shown (green, weak hydrogen bonds in blue).
Figure 5. Compound YH-53 docked to SARS-CoV2 3CL Pro (pdb 7E19). The docked ligand is represented in green, while the co-crystallized ligand is represented in dark gray. Hydrogen bond contacts are shown (green, weak hydrogen bonds in blue).
We can see that the catalytic cysteine, Cys145, makes a covalent bond with the ketone adjacent to the benzothiazole. Furthermore, despite the very flexible nature of this pseudopeptide, the docking pose is similar to the co-crystallized ligand (RMSD = 1.9 Å). The stabilizing hydrogen bonds are consistent between the two poses, highlighting the accuracy of our covalent docking.
An example of the pentafluorobenzene sulfonamide warhead
The PFBS warhead is unique from the aldehyde and activated ketone warheads in that a fluoride ion is released upon reaction. The nucleophilic residue attacks the para-fluorine of the pentafluorobenzene. One example of this covalent warhead in action is SH-4-54, an inhibitor of the Signal Transducer and Activator of Transcription 3 (STAT3) protein implicated in cancer.7-8 Although there is currently no co-crystal structure, 19F NMR studies suggest that the ligand is binding covalently due to NMR observation of a formed fluoride ion. 8-9 There is also evidence that the para-carbon of PFBS binds to the thiol of glutathione.8 Previously, this compound and its analogues have been docked non-covalently to the SH2 domain of STAT3,10-11 but considering the evidence of covalent binding, Flare can be used to predict how this compound might bind covalently to the cysteine in the active site. Figure 6 shows both covalent and non-covalent potential binding modes of SH-4-54.
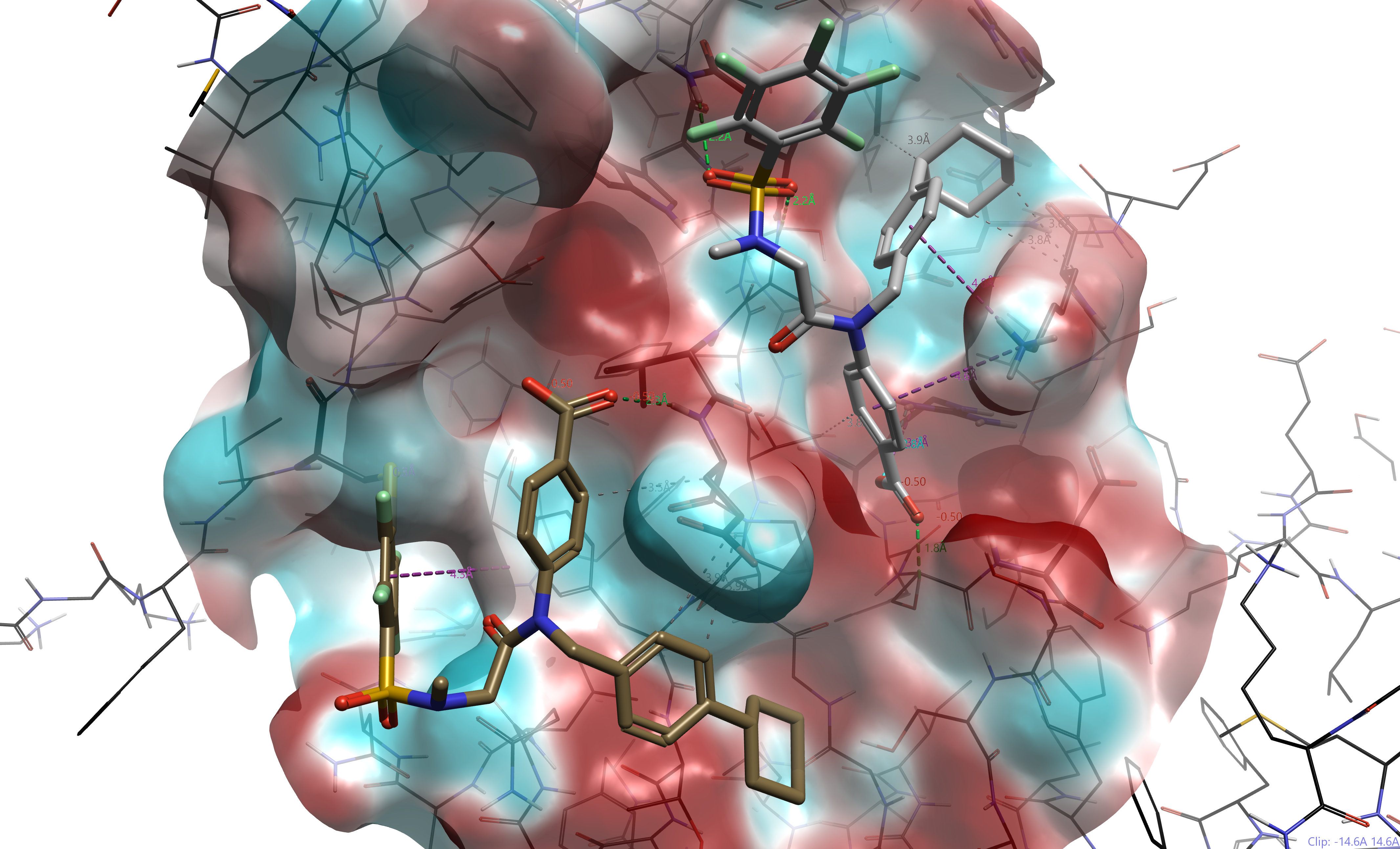 Figure 6. Docking SH-4-54 to STAT3 SH2 domain (pdb 1BG1). Brown: SH-4-54 covalently docked; Gray: SH-4-54 non-covalently docked. Although shown together, both poses were generated independently of one another.
Figure 6. Docking SH-4-54 to STAT3 SH2 domain (pdb 1BG1). Brown: SH-4-54 covalently docked; Gray: SH-4-54 non-covalently docked. Although shown together, both poses were generated independently of one another.
Docking SH-4-54 covalently, we see that the resultant tetrafluorobenzene participates in aromatic interactions with a nearby Tyr640 (shown by the purple dotted line), while the carboxylic acid participates in hydrogen bonding with a backbone NH from Glu638. Non-covalent (or Normal) docking reveals many more interactions of SH-4-54 in the binding site, including several hydrogen bonds and aromatic interactions. This binding mode is similar to one proposed by Page et al.11 In Figure 6, seeing the poses side by side, we see that the ligands are binding quite far from one another (RMSD = 15.4 Å). Unfortunately, despite the 19F NMR evidence of covalent binding, further experimentation is required to elucidate the true binding mode of SH-4-54.
Can custom warheads be used?
Yes, Flare is very customizable! Our Scientists and Support staff are more than happy to help you incorporate your custom covalent warheads. In the Flare Preferences (General tab), you are also able to upload your own .pat file which defines the compatible warheads and how they react.
If you foresee an issue with sharing IP, our staff can guide you in implementing your designs while keeping your data private to you.
You may also indicate the preferred warhead in the advanced covalent docking options (click the ‘Show Options’ button in the ‘Covalent Docking Calculation’ panel). For example, if your ligand covalently binds through its aldehyde, but the ligand also contains a higher-priority nitrile (Figure 3), you can input the SMARTS pattern corresponding to an aldehyde: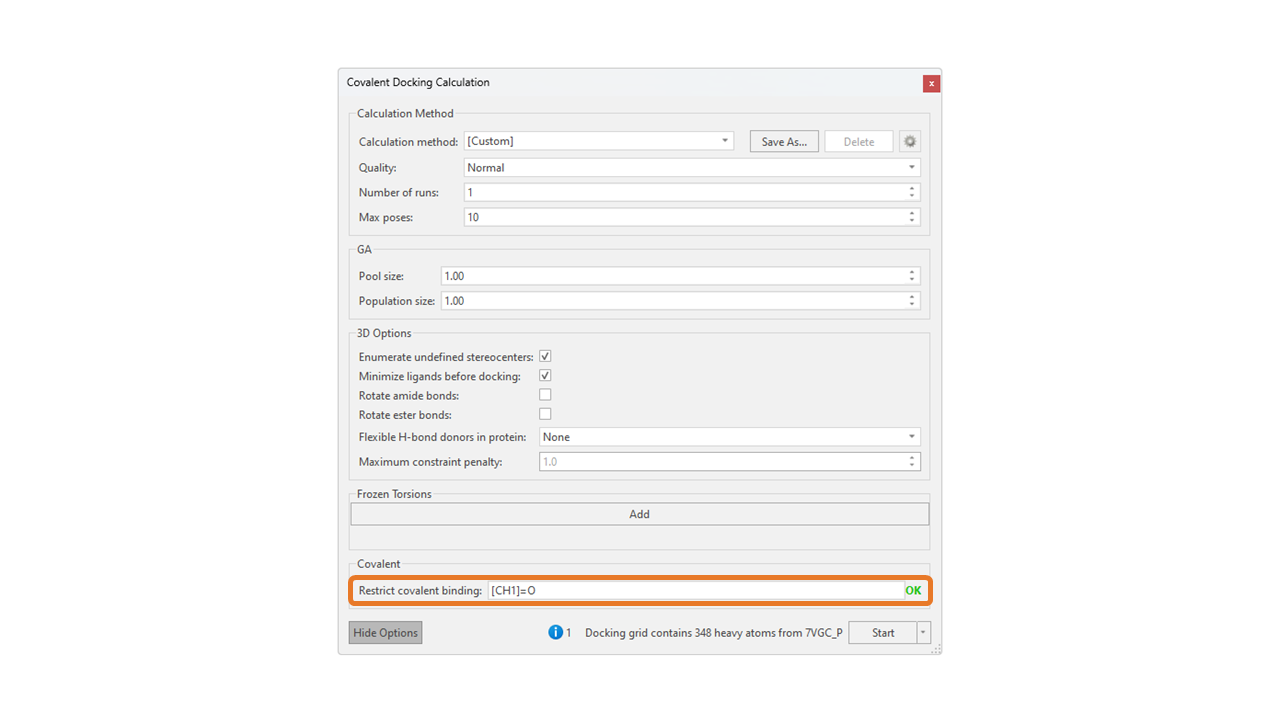 Figure 7. Advanced covalent docking options. The ‘Covalent’ section allows the user to input the SMARTS pattern of the prioritized warhead.
Figure 7. Advanced covalent docking options. The ‘Covalent’ section allows the user to input the SMARTS pattern of the prioritized warhead.
For any existing Flare customers who have questions or require further assistance in using warheads for Covalent Docking within Flare, please e-mail Cresset Support. If you’d like to explore the platform and its full range of CADD features in more depth, then please contact us today to request a free evaluation of Flare.
References
- Boike, L., Henning, N.J. & Nomura, D.K. Advances in covalent drug discovery. Nat Rev Drug Discov 2022. https://doi.org/10.1038/s41573-022-00542-z
- Singh, J., Petter, R., Baillie, T. et al. The resurgence of covalent drugs. Nat Rev Drug Discov 2011, 10, 307–317. https://doi.org/10.1038/nrd3410
- Huang, P., Lv, A., Yan, Q. et al. The structure and molecular dynamics of prolyl oligopeptidase from Microbulbifer arenaceous provide insights into catalytic and regulatory mechanisms. Acta Cryst 2022. D78, 735–751. https://doi.org/10.1107/S2059798322004247
- Sheppard, G. S., Florjancic, A. S. et al. Aryl ketones as novel replacements for the C-terminal amide bond of succinyl hydroxamate MMP inhibitors. Bioorg Med Chem Lett 1998, 8(22), 3251–3256. https://doi.org/10.1016/S0960-894X(98)00597-6
- Daum, S., Erdmann, F. et al. Aryl Indanyl Ketones: Efficient Inhibitors of the Human Peptidyl Prolyl cis/trans Isomerase Pin1. Angew Chem Int Ed 2006, 45(44), 7454–7458. https://doi.org/10.1002/anie.200601569
- Konno, S., Kobayashi, K. et al. 3CL Protease Inhibitors with an Electrophilic Arylketone Moiety as Anti-SARS-CoV-2 Agents. J Med Chem 2022, 65, 4, 2926–2939. doi.org/10.1021/acs.jmedchem.1c00665
- Linher-Melville, K., Nashed, M. G., et al. Chronic Inhibition of STAT3/STAT5 in Treatment-Resistant Human Breast Cancer Cell Subtypes: Convergence on the ROS/SUMO Pathway and Its Effects on xCT Expression and System xc- Activity. PLoS One 2016, 11(8), e0161202. doi.org/10.1371/journal.pone.0161202
- Ali, A., Gómez-Biagi, R., et al. Disarming an electrophilic warhead: Retaining potency in Tyrosine Kinase Inhibitor (TKI)-resistant CML lines, while circumventing pharmacokinetic liabilities. ChemMedChem. 2016 Apr 19; 11(8): 850–861. doi.org/10.1002/cmdc.201600021
- Wingelhofer, B., Maurer, B. et al. Pharmacologic inhibition of STAT5 in acute myeloid leukemia. Leukemia 2018, 32(5), 1135–1146. doi.org/10.1038/s41375-017-0005-9
- Huang, Q., Zhong, Y. et al. Structure-based discovery of potent and selective small-molecule inhibitors targeting signal transducer and activator of transcription 3 (STAT3). Eur J Med Chem 2021, 221, 113525. doi.org/10.1016/j.ejmech.2021.113525
- Page, B. D. C., Croucher, D. C. et al. Inhibiting Aberrant Signal Transducer and Activator of Transcription Protein Activation with Tetrapodal, Small Molecule Src Homology 2 Domain Binders: Promising Agents against Multiple Myeloma. J Med Chem 2013, 56, 7190−7200. dx.doi.org/10.1021/jm3017255